
-
Tatyana Igumenova
- Professor, Biochemistry and Biophysics
- Focus Area: signal transduction, membrane proteins, lipid membranes, lipid-activated kinases, AGC kinases, protein structure and dynamics, protein kinase C, synaptotagmin I, lipid-binding domains, xenobiotic metal ions, Pb(II) and Cd(II) toxicity, NMR
- Office:
- NMR N118A
- Email:
- [email protected]
- Phone:
- 979-314-8310 Lab phone: 979-314-8311
- Graduate Education
- Ph.D. Columbia University (2003)
- Postdoc. University of Pennsylvania (2003-2005)
- Postdoc. Columbia University (2005-2007)
Areas of Expertise
Lipid signaling and its dysregulation in disease, membrane proteins, integrative structural biology, Nuclear Magnetic Resonance (NMR) Spectroscopy, X-ray crystallography, atomistic molecular dynamics simulations.
Professional Summary
Dysregulation of signal transduction networks is the root cause of many human diseases such as cancer, cardiac disease, and intellectual disabilities. The major theme of my laboratory focuses on understanding the structural basis of signal transduction on membrane surfaces. Specifically, we study protein systems that regulate key signaling events such as lipid- and calcium-dependent phosphorylation, neurotransmission, and temporal control of protein conformational changes on biologically relevant timescales. Many of these events involve reversible association of signaling proteins with membranes in response to different stimuli, and we are particularly interested in deciphering how multi-modular protein architectures support these signaling functions. To address those questions, we employ an integrative structural biology approach that involves advanced solution Nuclear Magnetic Resonance (NMR) spectroscopy, X-ray crystallography, and atomistic molecular dynamics simulations of protein-membrane systems.
Protein Kinase C: structure, regulation, and opportunities for drug design
Protein Kinases C (PKC) defines a family of lipid-activated kinases that play vital roles in the phosphoinositide-signaling pathway. Classical views as to how PKCs function were recently overturned by the paradigm-shifting discovery that PKC – long considered to function exclusively as a tumor promoter – phenotypes as a tumor suppressor in 8% of mutations associated with human cancers. Those surprising discoveries underscore the key challenges in the field – a need for the availability of both activators and inhibitors of PKC function, and the ability to target the protein in an isoform-specific manner. Interference with the PKC function by direct targeting of its diacylglycerol-sensitive C1 domains is a research direction that promises launch of fresh pharmaceutical approaches for cancer therapies, Alzheimer’s disease, and HIV/AIDS. To that end, rational design of potent C1 modulators requires detailed information regarding: (i) functional properties of the ligand binding site; (ii) key residues that participate in ligand recognition and capture; (iii) role of binding site dynamics in ligand interactions; and (iv) role of other signaling lipids in the formation of C1-ligand complexes. We are applying X-ray and NMR structural approaches, in combination with MD simulations, to determine the structural basis of C1-ligand-membrane interactions for a chemically diverse set of exogenous PKC agonists.
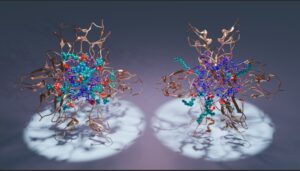
Assembly of C1 domains (gold) around micelles composed of diacylglycerol (blue) and dodecylphosphocholine (cyan).
Katti S.S., Krieger I.V., Ann J., Lee J., Sacchettini J.C., Igumenova T.I. (2022) Structural anatomy of Protein Kinase C C1 domain interactions with DAG and other agonists. Nature Communications 13:2695, doi.org/10.1038/s41467-022-30389-2.
Lipid transfer proteins as antifungal targets
Phosphatidylinositol/phosphatidylcholine exchange proteins (PITPs) are key players in the inositol lipid-based phosphoinositide signaling pathways. Through their lipid-exchange function, PITPs integrate lipid metabolism with phosphoinositide signaling. These do so by executing a heterotypic lipid exchange reaction of phosphatidylcholine for phosphatidylinositol which, in turn, stimulates the activities of phosphatidylinositol 4-OH kinases to promote specific phosphatidylinositol-4-phosphate signaling events related to membrane trafficking and cell cycle regulation. Studies conducted in S. cerevisiae demonstrate that Sec14 PITP is an attractive target for the development of anti-fungal therapies that interfere with phosphoinositide signaling without targeting the highly conserved lipid kinases themselves. This project seeks to develop and characterize next generation inhibitors with improved potencies and specificities tailored to targeting of the Sec14 PITPs of virulent fungi – particularly of Candida species such as albicans and auris.
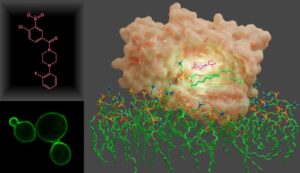
Small-molecule inhibitor (pink) complexed to Sec14 from S. cerevisiae.
Chen, X.-R., Poudel, L., Hong, Z., Johnen, P., Katti, S., Tripathi, A., Nile, A.H., Green, S., Khan, D., Schaaf, G., Bono, F., Bankaitis, V.A., and Igumenova, T.I. (2023) Mechanisms by Which Small Molecules of Diverse Chemotypes Arrest Sec14 Lipid Transfer Activity. Journal of Biological Chemistry, 299(2), 102861. Selected as “Recommended Reads”.
Xenobiotic metal ions: in-cell detection and structural basis of toxicity
“Xenobiotic” is defined as a “chemical … that is foreign to a living organism”. Metal ions that do not have a nutritive value belong to this category; some examples include rare earth metals and potent environmental toxins, such as Cd, Pb, As, and Hg. We discovered that some xenobiotic metal ions, such as Pb2+ and the lanthanide series, Ln3+, have unique physicochemical properties to their interactions with oxygen-rich Ca2+-binding sites of C2 domains, a class of Ca2+-dependent peripheral membrane modules found in >100 signaling proteins. We are exploiting these unique properties to address long-standing mechanistic questions about metal-ion mediated signal transduction and Cd/Pb toxicity. In addition to the in-vitro work, we are applying in-cell NMR approaches to probe metal ion binding to their target proteins in the crowded intracellular environments. Characterization of protein interactions with toxic metals in cellular contexts is required for reliable identification of physiologically relevant targets, development of strategies for bioremediation, and creation or refinement of protein-based sensors.

Schematic representation of in-cell NMR experiments (left panel) and comparsion of lysate and in-cell NMR spectra of heavy-metal ion sensor protein a3DIV (right panel).
Katti, S.S. and Igumenova, T.I. (2023) Protein-cadmium interactions in crowded biomolecular environments probed by in-cell and lysate NMR spectroscopy. doi: https://doi.org/10.1101/2023.11.03.565546.
Regulation of kinases by Pin1, a peptidyl-prolyl isomerase
Pin1 is a peptidyl-prolyl isomerase that catalyzes cis-trans isomerization of pThr/pSer-Pro bonds in its target proteins. Moreover, Pin1 regulates the activities of proteins involved in cancer initiation and progression, and Pin1 is overexpressed in various cancers. In-cell studies show that Pin1 interacts with, and regulates, the stabilities of at least six AGC kinases: PKC, Akt, S6K, SGK, GRK2, and LATS. Recent work on the Pin1-PKC interactions from our laboratory revealed that, contrary to prevailing models, Pin1 controls PKC activity via sequestration of the disordered C-terminal tail of the kinase. This novel mechanism involves a bivalent recognition of the two conserved phosphorylated PKC motifs by Pin1. This discovery is paradigm-shifting in that it provides the first example of a non-canonical and non-catalytic Pin1 activity in targeting hydrophobic motifs of AGC kinases as a mode of regulation of these key enzymes. Our current research efforts in this direction are focused towards: (i) developing sensitive methods for detection of Pin1 interactions with therapeutic agents, and (ii) establishing molecular mechanisms of Pin1 dysregulation in disease by understanding the functional derangements caused the cancer-related Pin1 mutations.

Structure of Pin1 complexed to the C-terminal region of PKCbII (top panel) and the proposed mechanism of PKCbII regulation by Pin1 (bottom panel). Chen, X.-R., Dixit, K., Yang, Y., McDermott, M.I., Imam, H. T., Bankaitis, V.A., and Igumenova T.I. (2023) A novel bivalent interaction mode underlies a non-catalytic mechanism for Pin1-mediated Protein Kinase C regulation”. doi: https://doi.org/10.1101/2023.09.18.558341.
Publications
- View publications on PubMed